11/17/2021
PITTSBURGH – By coupling machine learning with whole genome sequencing, University of Pittsburgh School of Medicine and Carnegie Mellon University scientists greatly improved the quick detection of infectious disease outbreaks within a hospital setting over traditional methods for tracking outbreaks.
The results, published today in the journal Clinical Infectious Diseases, indicate a way for health systems to identify and then stop hospital-based infectious disease outbreaks in their tracks, cutting costs and saving lives.
“The current method used by hospitals to find and stop infectious disease transmission among patients is antiquated. These practices haven’t changed significantly in over a century,” said senior author Lee Harrison, M.D., professor of infectious diseases at Pitt’s School of Medicine and epidemiology at Pitt’s Graduate School of Public Health. “Our process detects important outbreaks that would otherwise fly under the radar of traditional infection prevention monitoring.”
The Enhanced Detection System for Healthcare-Associated Transmission (EDS-HAT) couples the recent development of affordable genomic sequencing with computer algorithms connected to the vast trove of data in electronic health records. When the sequencing detects that any two or more patients in a hospital have near-identical strains of an infection, machine learning quickly mines those patients’ electronic health records for commonalities – whether that be close proximity of hospital beds, a procedure using the same equipment or a shared health care provider – alerting infection preventionists to investigate and halt further transmission.
 Ordinarily, this process requires clinicians to notice that two or more patients have a similar infection and alert their infection prevention team, which can then review patient records to attempt to find how the infection was transmitted.
Ordinarily, this process requires clinicians to notice that two or more patients have a similar infection and alert their infection prevention team, which can then review patient records to attempt to find how the infection was transmitted.
“This is an incredibly labor-intensive process that is often dependent upon busy health care workers noticing a shared infection between patients to begin with,” said lead author Alexander Sundermann, M.P.H., C.I.C., F.A.P.I.C., clinical research coordinator and doctoral student at Pitt Public Health. “That might work if patients are in the same unit of a hospital, but if those patients are in different units with different health care teams and the only shared link was a visit to a procedure room, the chances of that outbreak being detected before other patients are infected falls dramatically.”
From November 2016 to November 2018, UPMC Presbyterian Hospital ran EDS-HAT with a six-month lag for a few select infectious pathogens often associated with health care-acquired infections nationwide, while continuing with real-time, traditional infection prevention methods. The team then investigated how well EDS-HAT performed.
EDS-HAT detected 99 clusters of similar infections in that two-year period and identified at least one potential transmission route in 65.7% of those clusters. During the same period, infection prevention used whole genome sequencing to aid in the investigation of 15 suspected outbreaks, two of which revealed genetically related infections.
If EDS-HAT had been running in real-time, the team estimates as many as 63 transmissions of an infectious disease from one patient to another could have been prevented. It also would have saved the hospital as much as $692,500.
 In one case-study, EDS-HAT found an outbreak of vancomycin-resistant Enterococcus faecium that it traced to an interventional radiology procedure involving injection of sterile contrast that was being performed according to manufacturer instructions. Due to EDS-HAT detecting the outbreak, UPMC alerted the manufacturer to the instructions that led to faulty sterilization practices.
In one case-study, EDS-HAT found an outbreak of vancomycin-resistant Enterococcus faecium that it traced to an interventional radiology procedure involving injection of sterile contrast that was being performed according to manufacturer instructions. Due to EDS-HAT detecting the outbreak, UPMC alerted the manufacturer to the instructions that led to faulty sterilization practices.
“In that case, EDS-HAT connected the dots between seemingly unconnected patient infections occurring in different hospital units, stopping that outbreak but also potentially preventing similar outbreaks at other hospitals,” Harrison said. “That example encapsulates the value of EDS-HAT.”
UPMC plans to introduce EDS-HAT in real-time at UPMC Presbyterian Hospital and expects this innovation to benefit other infection prevention and control programs in the future. And the original EDS-HAT, which primarily focused on drug-resistant bacterial pathogens, will soon be expanding to incorporate sequencing of respiratory viruses, including COVID-19.
Additional authors on this research are Jieshi Chen, M.S., James K. Miller, Ph.D., and Artur Dubrawski, Ph.D., all of Carnegie Mellon University; and Praveen Kumar, B.Tech., Ashley M. Ayres, Shu-Ting Cho, M.S., Chinelo Ezeonwuka, M.Sc., Marissa P. Griffith, Mustapha M. Mustapha, Ph.D., A. William Pasculle, Sc.D., Melissa I. Saul, M.S., Kathleen A. Shutt, M.S., Vatsala Srinivasa, M.P.H., Kady Waggle, Daniel J. Snyder, M.Sc., Vaughn S. Cooper, Ph.D., Daria Van Tyne, Ph.D., Graham M. Snyder, M.D., Jane W. Marsh, Ph.D., and Mark S. Roberts, M.D., all of Pitt or UPMC, or both.
This study was funded in part by National Institute of Allergy and Infectious Diseases grants R21 AI109459 and R01 AI127472.
PHOTO INFO: (click images for high-res versions)
PHOTO INFO: (click images for high-res versions)
CREDIT BOTH: Nathan Langer/UPMC
Top:
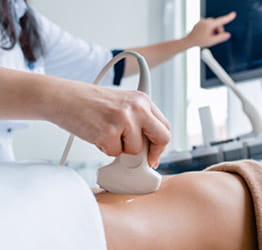
CAPTION: Dr. Lee Harrison and Alex Sundermann load samples for genomic sequencing
Bottom:
CAPTION: Dr. Lee Harrison points out to Alex Sundermann a potential outbreak detected by EDS-HAT