12/13/2018
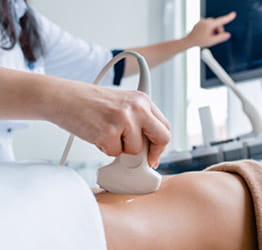
PITTSBURGH – Using statistical models to analyze data from nearly 2,000 families with an autistic child, a multi-institute research team discovered tens of thousands of rare mutations in noncoding DNA sequences and assessed if these contribute to autism spectrum disorder.
Published today in the journal Science, the study is the largest to date for whole-genome sequencing in autism. It included families comprised of both biological parents, a child affected with autism and an unaffected sibling.
Scientists representing University of Pittsburgh School of Medicine, Carnegie Mellon University, University of California, San Francisco (UCSF), Massachusetts General Hospital, Harvard Medical School and the Broad Institute led the research team.
The study is one of 13 released in the first round of results to emerge from the National Institute of Mental Health’s PsychENCODE Consortium — a nationwide research effort that seeks to decipher how noncoding DNA, often referred to as the ‘dark matter’ of the human genome, contributes to psychiatric diseases such as autism, bipolar disorder and schizophrenia.
Over the past decade, scientists have identified dozens of genes associated with autism by studying so-called “de novo” mutations — newly arising changes to the genome found in children but not their parents. To date, most de novo mutations linked to autism have been found in protein-coding genes. It has proven far more difficult for scientists to identify autism-associated mutations in noncoding regions of the genome.
“We were particularly interested in the elements of the genome that regulate when, where and to what degree genes are transcribed. Understanding this noncoding sequence could provide insights into a variety of human disorders,” said Bernie Devlin, Ph.D., corresponding author and professor of psychiatry at the Pitt School of Medicine.
“Protein-coding genes clearly play an important role in human disorders like autism, yet their expression is regulated by the ‘noncoding’ genome, which covers the remaining 98.5 percent of the genome and remains somewhat mysterious,” said Carnegie Mellon’s Kathryn Roeder, Ph.D., corresponding author and UPMC Professor of Statistics and Life Sciences. “Because the genome comprises 3 billion nucleotides, identifying which portions of the noncoding genome, when mutated, enhance the risk of autism is as challenging as looking for a needle in a haystack.”
Using a novel bioinformatics framework, the researchers were able to compress the search from billions of nucleotides to tens of thousands of functional categories that potentially contribute to autism. Working with these categories, they used machine learning tools to build statistical models to predict autism risk from a subset of the families in the study. They then applied this model to an independent set of families and successfully predicted patterns of risk in the noncoding genome.
Though rare de novo mutations were found in many noncoding regions of the genome, the strongest signals arose from promoters — noncoding DNA sequences that control gene transcription. These risk-conferring promoters were most often located far from the genes under their control. They also were found to be largely conserved across species, suggesting that any rare mutations that might arise in these promoters are more likely to disrupt normal biology.
“For years, scientists have used genome-wide studies to find common variants that confer disease risk. Our group has now focused on creating a computational framework that’s capable of finding rare, high-impact variants associated with a human disorder, looking across all the noncoding regions of the genome,” said Stephan Sanders, Ph.D., corresponding author and professor of psychiatry at the UCSF Weill Institute for Neurosciences and Institute for Human Genetics.
The team’s findings have practical implications for future research on model organisms, like mice, as attempts are made to move toward genetically informed therapies for autism. But the value of studying the noncoding genome extends well beyond autism.
“We are just scratching the surface of what there is to learn about noncoding regulatory variation in human disease, and the new methods this team has developed will catalyze an important step forward into larger and more comprehensive studies,” said Michael Talkowski, Ph.D., of Massachusetts General Hospital, Harvard Medical School and the Broad Institute, who also served as corresponding author on the study.
Lead authors on the paper are Joon-Yong An, Ph.D, and Donna Werling, Ph.D, of the UCSF Weill Institute for Neurosciences; and Kevin Lin, Ph.D., and Lingxue Zhu, Ph.D., of CMU’s Department of Statistics and Data Science.
The National Institutes of Health, the Simons Foundation Autism Research Initiative and the Broad Institute’s Stanley Center for Psychiatric Research provided funding for this research.